Dr. Rick Lifton On Genes, Genomes, and The Future Of Medicine

Editor’s Note: In honor of SSRP ’19 beginning this year, we’ve decided to rerelease an interview conducted by Ella Epstein (SSRP ’16-’17) with The Rockefeller University President, Dr. Lifton.
Over the [2017] summer, Rockefeller University president Dr. Lifton gave a lecture on the impact of genetics on human health and disease. Interested to learn more, I reached out to Dr. Lifton to ask him more about his work.
What does your lab do?
“My lab uses human genetics and genomics to identify mutations that contribute to disease. A number of years ago, we developed methods for rapidly and inexpensively sequencing all 20,000 protein-coding genes in the human genome, and have been applying this technology to look for diseases that are caused by rare mutations with large effects that would not be discoverable with prior technologies. The consequence of this work has been the realization that a large fraction, likely a majority of the 20,000 protein coding genes, will have consequences for health and disease when they are mutated. Given that we know the consequence of mutation of less than 4,000 of these genes, it indicates that an awful lot of work lies ahead.”
How do you differentiate between genetic risk factors and harmless mutations that occur over time? How do you pinpoint specific mutations that cause disease versus any other mutation that might have occurred in the same person?
“The genes that we can most readily translate into new targets for therapy are the mutations that have a large effect on health risks. If you find mutations that are clearly driving health and disease, these directly identify pathways that might be manipulated for health benefit, both in patients with mutations but also in the general population. There are examples where the rare mutations point to pathways that could be manipulated for health benefit in everyone. For instance, we know that high LDL cholesterol causes heart attacks, and that knowledge came from understanding the biology underlying a very rare disease. However, we now treat tens of millions of Americans who, although they do not have such mutations, benefit from medications that lower LDL cholesterol levels.”
How did you get the idea for your research?
“I got a background in Drosophila genetics, and at that time we were starting to apply the that if we knew that there were mutations that were causing a trait, we might be able to find the genetic mutation that underlies that trait. That was back in the late 1970s, when I was a graduate student. I was an MD – PHD student at Stanford, so I finished my PHD, finished medical school, and went off to clinical training at Harvard and did my clinical training in internal medicine, and came out of that experience with the idea that we could now begin to apply the same kinds of approaches that we were using in fruit flies to understanding human disease. Hypertension looked like the kind of important problem that this technology would be well suited to, because it affected so many people that nobody could come to conclusions where the primary abnormalities that were causing the big disease might lie. So, we thought if we could find single mutations that completely disrupted the normal regulation of the system, pushing the blood pressure either to one extreme or the other, that this would begin to identify pathways that might be regulating blood pressure, not just in rare individuals, but in everybody. Hypertension affects a billion people worldwide and it’s a major contributor to deaths from cardiovascular disease. And because we can only study high blood pressure in the context of a living human being or an animal, it’s very difficult to figure out from physiology alone where the primary abnormalities that are driving high and low blood pressure lie. So we took a very simple genetic approach to this complicated problem and looked for people with the highest of blood pressures around the world, screening them for individual single gene mutations that could be driving blood pressure to the extremes. And we did that by scouring the medical literature, looking for case reports of unusual patients with extremely high and low blood pressure, making contact with people in specialty clinics who see these patients from around the world. Over the years, we have collected thousands of samples from patients with these extreme forms of high and low blood pressure, and from that set, identified mutations in about a dozen genes that drove the blood pressure to the extreme high end of the distribution, and another dozen that drove blood pressure to the extremely low end of the distribution. And the importance of the work is that all of the genes converge on a single common pathway that regulates how the kidneys handle salt. So, if your kidneys don’t reabsorb more salt, blood pressure goes down; if they reabsorb it, blood pressure goes up.”
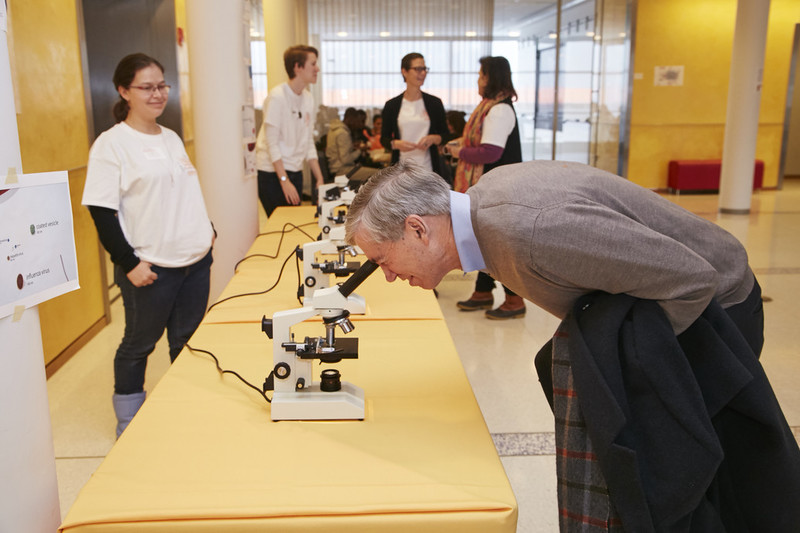

Dr. Lifton at Talking Science, an annual event of interactive lectures and activities at Rockefeller for high schoolers.
It seems like you were looking to solve the issue of doctors not addressing the basic science in addressing these problems. What’s the alternative? What else would you do, besides basic science?
“We have a long history in medicine of doing our best in treating patients when we understand the fundamental causes of a disease. In general, when we don’t understand the cause of disease, we just try to empirically try things to see if something will work. And sometimes we hit upon effective treatments, but usually they are not as effective as they would be if we really understood the biology underlying the diseases we are trying to treat. This doesn’t always work perfectly. Sometimes we find the specific causes of disease and still struggle to find effective ways to prevent or treat disease. But it defines the playing field that we have to operate in. If we want to mitigate the effect of disease, we have to understand its basic causation.”
How do you see your research in relation to the developing field of personalized medicine?
“I think there are two broad approaches that will apply to health in the future. One might broadly think of these as the cardiovascular disease treatment model and the cancer therapeutic treatment model. In cardiovascular disease, we are finding the pathways that could be manipulated, allowing us to treat millions of people without knowing their personal genomes in great detail. And the reason for that is that many individuals who do not have single mutations that drive the pathogenesis of high LDL cholesterol or high blood pressure would still benefit from lowering their cholesterol or blood pressure. But by knowing the pathways that have come from studying rare patients, we can find effective ways to treat bigger groups of patients. Finding these rare mutations can identify the target to be used in the general population. That’s one model where understanding the basic biology can lead to medicines that can be broadly applied to the general population. In patients with cancer, knowing the specific mutations underlying disease are critically important in picking the right therapeutics. So, for example, if you have leukemia that is caused by mutations in Abelson kinase, or ABL, then you will respond extremely well to an inhibitor to that kinase. There is a drug called Glivec that is efficacious in patients who have that mutations. However, patients with leukemia who do not have mutation rarely benefit at all from that treatment. So this is a beautiful example of personalized medicine – here, knowing whether a patient does or does not have a specific mutation is critical in selecting that patient’s treatment methods. When we think about personalized medicine broadly, I think we will need to distinguish those diseases in which knowing the specific mutations in individual patients makes a critical difference in picking the right drug, and distinguishing those from cases where knowing the pathways that we can manipulate for health benefits are sufficient to be able to treat patients, without knowing much or anything about their genetics.”
Do you have any specific research-oriented goals at Rockefeller?
“We know there are 20,000 genes in a human genome. We know what happens if about 4,000 of those genes are mutated. This is now a bounded problem. We know there are 20,000 protein-coding genes and no more. There are no fundamental obstacles now to identifying the consequence of mutation in all 20,000 genes. And as a field, this a key part of the “to-do” list. Understanding the consequence of mutations of each of these genes will define their contribution to health and disease and also identify a wide range of potential therapeutic targets that can be manipulated for health benefit in the population. This is an important goal for advancing and understanding health and disease for the next decade.”